The available modules of the LC-MS workflow can be grouped into three categories: pre-processing, statistics, and annotation (Figure 2). The modules for statistical analysis can also be used with processed data (sample x variable tables) coming from NMR or other "omic" technologies.
 Detailed trainings are available online https://training.galaxyproject.org/training-material/topics/metabolomics/
Detailed trainings are available online https://training.galaxyproject.org/training-material/topics/metabolomics/
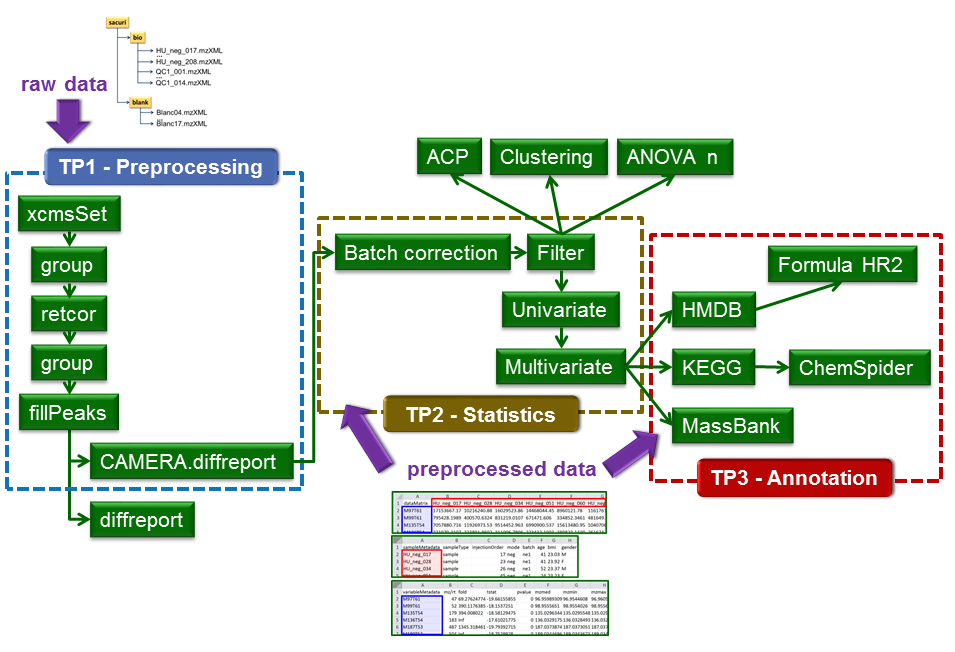
Figure 2: Available modules for LC-MS analysis. Raw data (in .mzXML or .cdf formats to be handled by XCMS) can be uploaded by using an FTP protocol. Pre-processed data (i.e. variables x samples data matrix and the two associated tables for samples and variables metadata) can also be uploaded directly via the Galaxy interface.
Modules can be selected and chained by the user via the workflow editor (Figure 3). A dataset (named 'sacuri') is provided as an example. It contains:
- 20 LC-MS spectra of individual human urine samples (high-resolution LTQ-Orbitrap spectrometer; negative ionization mode),
- 10 quality control (pooled) samples,
- and 6 blanks (water only).
The details of each analysis step are provided in the following sessions.
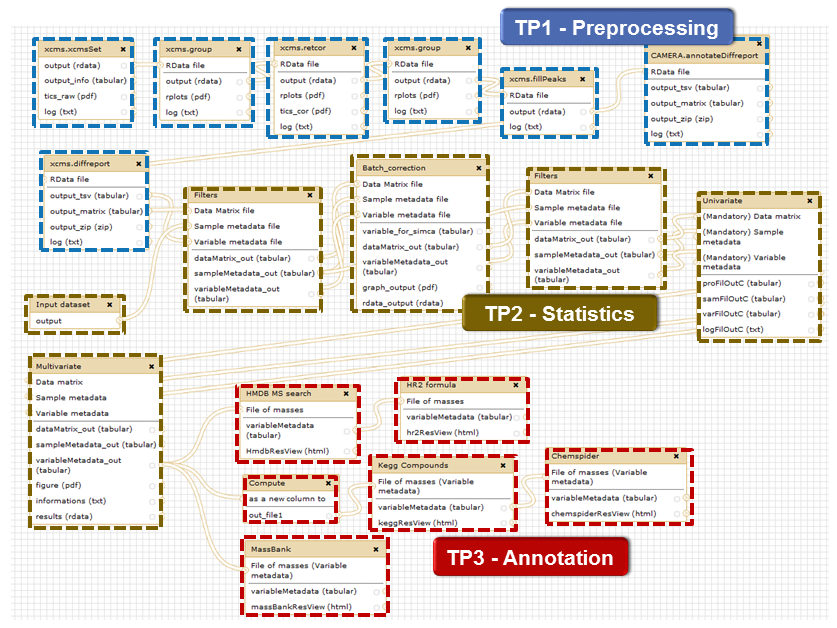
Figure 3: Example of a full LC-MS workflow chaining all available modules. Example sub-workflows for pre-processing, statistical analysis, and annotation are available on the W4M site.